Hi All,
So as I’ve discussed before I’ve been tagging microarray text annotations with ontology tags. I’ve also discussed how I use ROC curves to measure the accuracy of the predictions. I’ve finally finished all of the back-end work and now I’ve been getting results.
First, ROC curves:

Predction ability of MMB
These have AUCs of 0.94, 0.95, 0.87 for the Disease Type, Functional Anatomy and Cell Type respectively. I have hand-annotated a LARGE number of microarray records, using a method I’ll describe later in this post. Some details: I require that a GSM record has been annotated with AT LEAST 1 TP and 2 TNs to be included in the ROC calculations … This results in the following numbers of training datasets … By default I use 2/3rds of the data for training and 1/3rd for testing … also the implementation of the MaxEntropy classifier that I use cannot train with TN annotations, so they are only used for the “specificity” calculation:
- Disease Type: 1051 TPs and 96533 TNs
- Functional Anatomy: 523 TPs and 31017 TNs
- Cell Type: 677 TPs and 16489 TNs
While I am ECSTATIC about these results I think they are “suspiciously high”. Most of the NLP research that I’ve combed through has AUCs (or comparable measures) that are closer to 0.7-0.8 … granted, these studies were on POS tagging or “Named Entity Recognition” (gene name identification) and relied only on features from a single sentences. In my search to confirm or deny my prediction abilities I stumbled across something interesting which I’ll share here:
When I plot out a histogram of the predicted probabilites I notice an intersting side-effect of my annotation strategy:
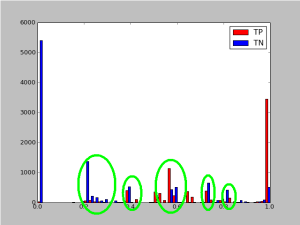
A histogram of the probilities of labeled tags
When I hand-curate the predictions I use the django-admin actions function. I’ve created a simple admin-action in the predictions admin interface which essentially creates an “agreement” or “disagreement” with the predicted tag:
A screen-shot of the admin-action website
I simply check-off the predictions I agree with and then use the admin-action to create a hand-annotation. When I sort by prediction-probability I get another bonus … It will group similar tags and similar descriptions together, this happens because the extracted features for many GSM records are identical so they have the same predicted tag with the same probability. Due to this grouping it is trivial for me to scan through the predictions … I can annotate ~500 GSM records in about 10 minutes.
This is what created the “blips” that are circled in this histogram. When I do my annotations I’ll usually scroll in the admin interface to a random probability. Then I’ll annotate ~2000 GSM records and then scroll to another probability.
After I noticed this I tried to annotate in a “truly” random method by having the admin interface shuffle the predictions it gave me. However, this DRASTICALLY reduced my annotation throughput … it took ~45 min to do 500 annotations (nearly 10-fold slower). This because the records weren’t grouped and I had to read each one more carefully, I couldn’t just skim it to ensure it was identical to the previous one.
If anyone has any suggestions for how to correct my curation bias yet maintain my throughput I’d greatly appreciate it.
Will
just FYI … the admin actions are actually “development features” of django and are not in the “stable release”, only the SVN version. So use at your own risk and YMMV.